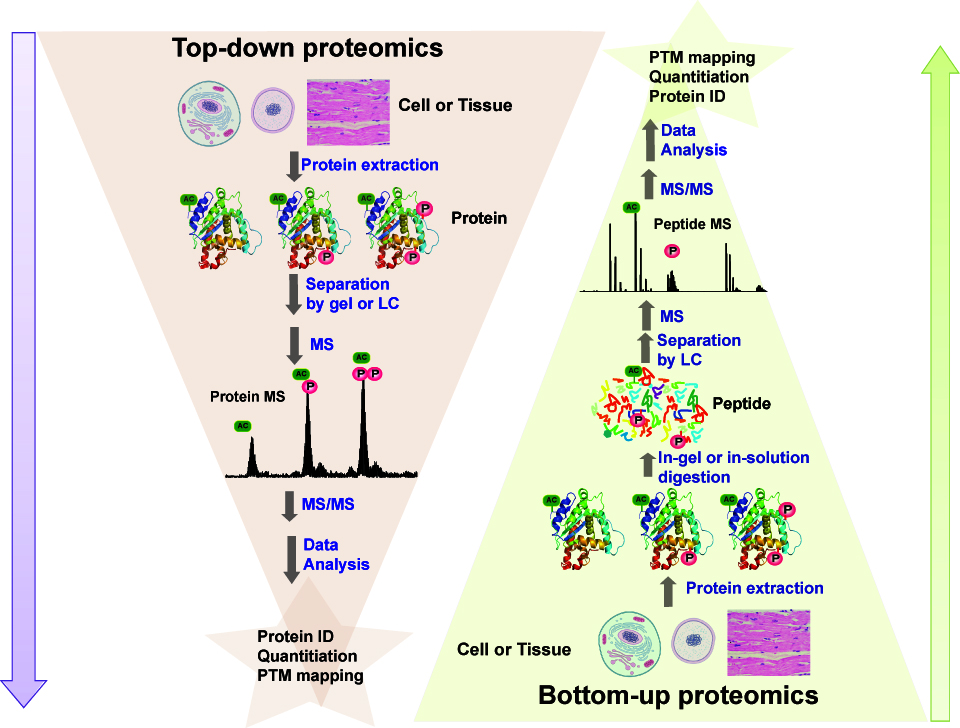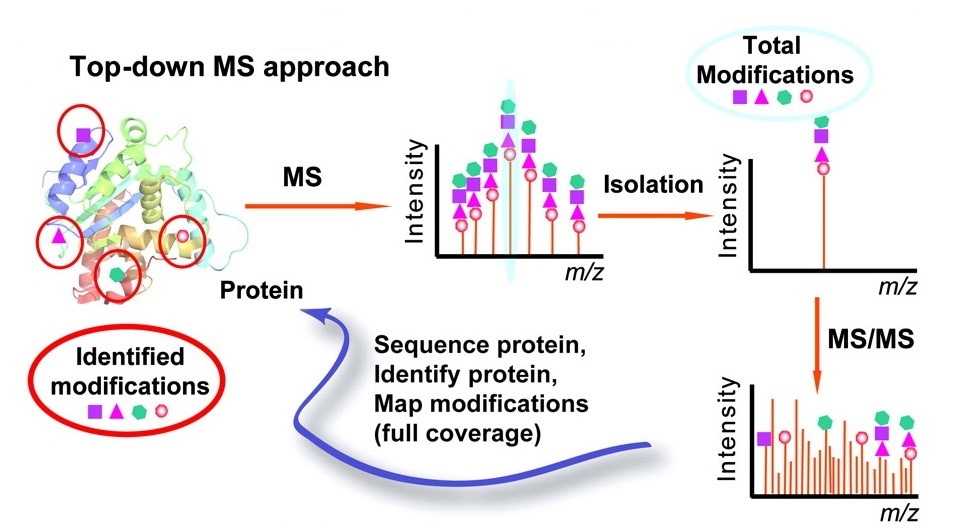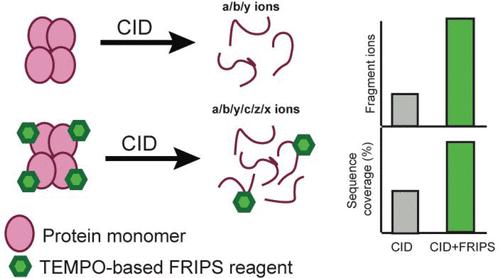
Free Radical-Based Sequencing for Native Top-Down Mass Spectrometry,Journal of the American Society for Mass Spectrometry - X-MOL

Free Radical-based Sequencing for Native Top-Down Mass Spectrometry | Analytical Chemistry | ChemRxiv | Cambridge Open Engage
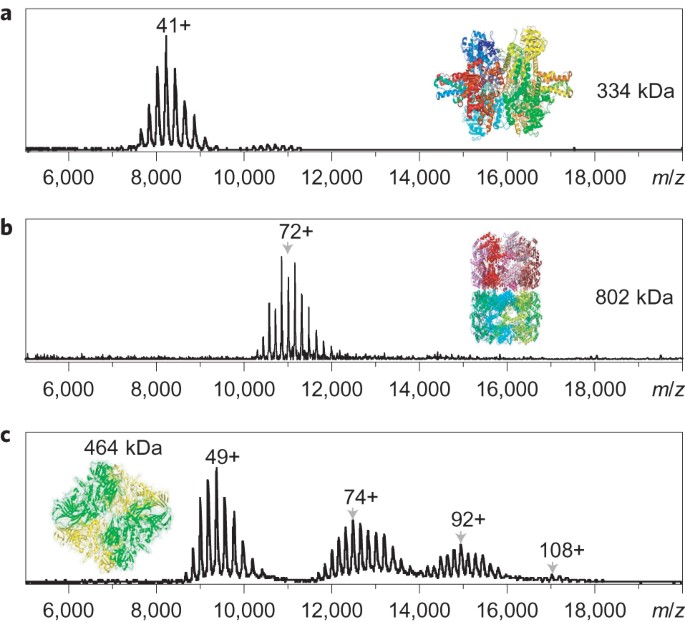
An integrated native mass spectrometry and top-down proteomics method that connects sequence to structure and function of macromolecular complexes | Nature Chemistry

Characterizing the top-down sequencing of protein ions prior to mobility separation in a timsTOF - Analyst (RSC Publishing)

MALDI Top-Down sequencing: calling N- and C-terminal protein sequences with high confidence and speed. - Abstract - Europe PMC

Top-down proteomics for the analysis of proteolytic events - Methods, applications and perspectives - ScienceDirect
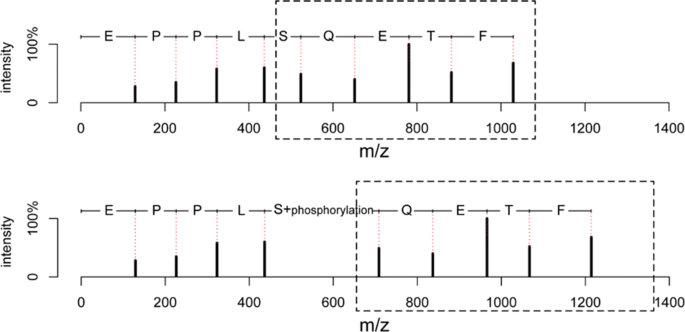
Evaluation of top-down mass spectral identification with homologous protein sequences | BMC Bioinformatics | Full Text

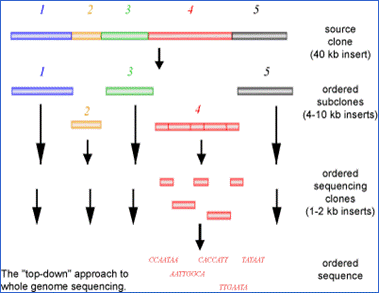
Difference Between Hierarchical and Whole Genome Shotgun Sequencing | Compare the Difference Between Similar Terms
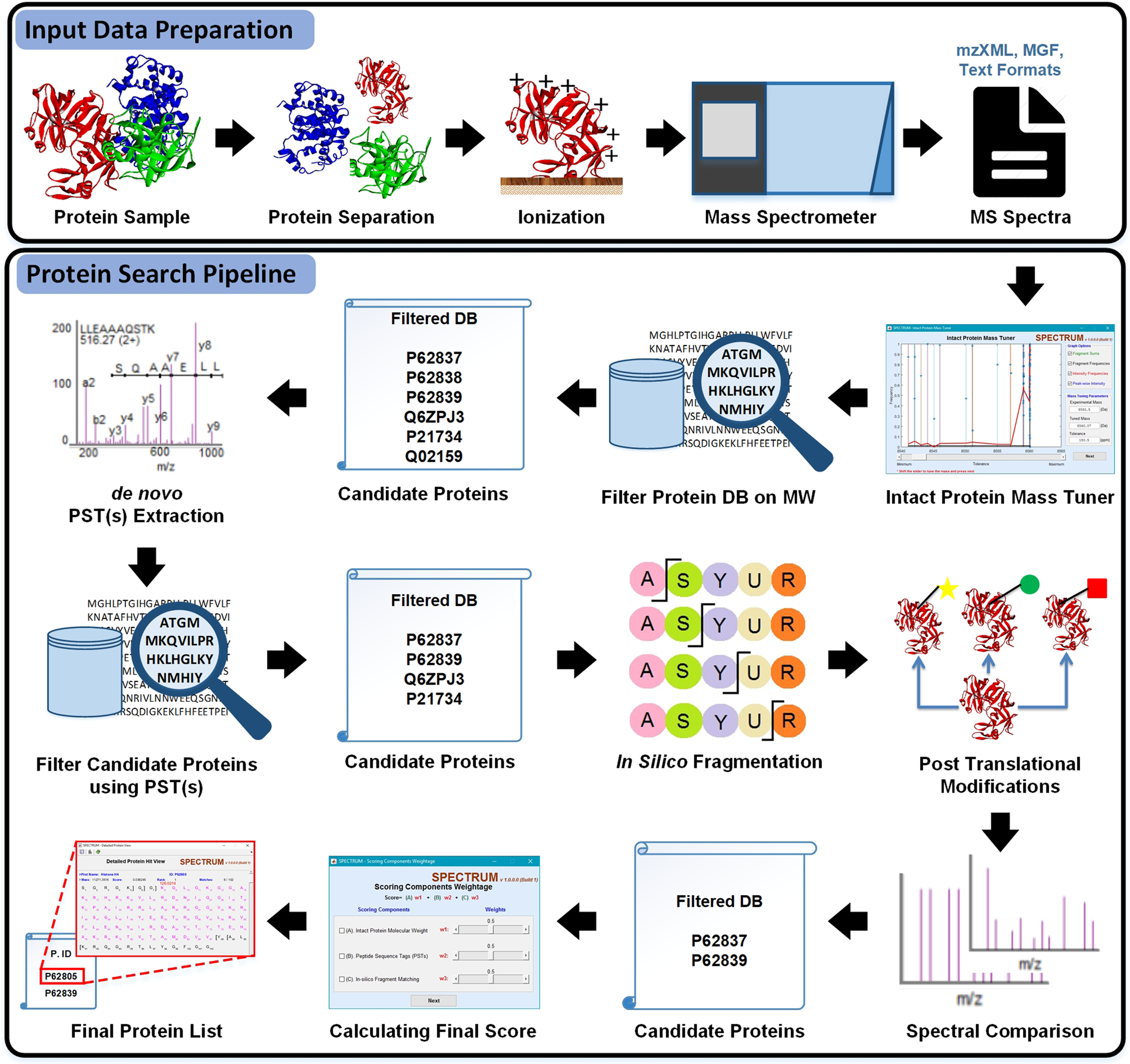
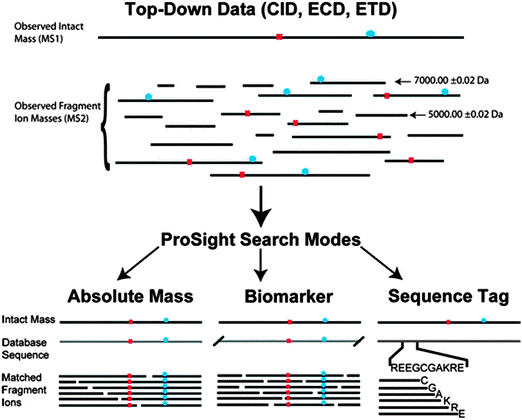
SPECTRUM – A MATLAB Toolbox for Proteoform Identification from Top-Down Proteomics Data | Scientific Reports

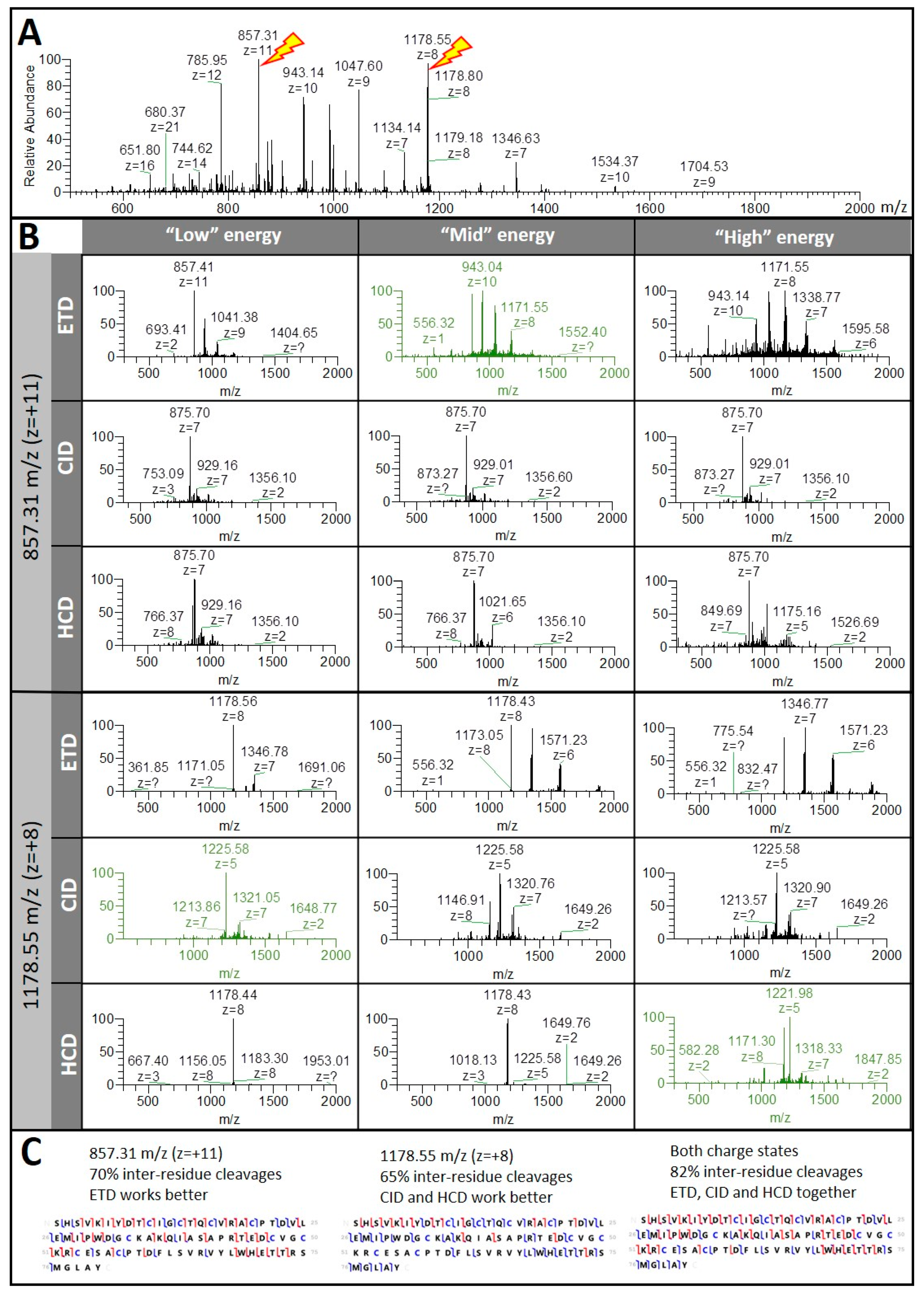
Maximizing Sequence Coverage in Top-Down Proteomics By Automated Multimodal Gas-Phase Protein Fragmentation | Analytical Chemistry